Benefiting from the Wealth of Personal Time-Series on Personal Devices
An ever-increasing amount of data is expected to
be generated at the individual side (e.g.,
wearables, smart-meters) (i) often as time-series
and (ii) stored on a personal device for being
analyzed and vizualized.
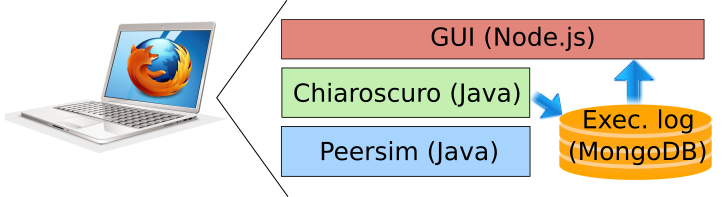
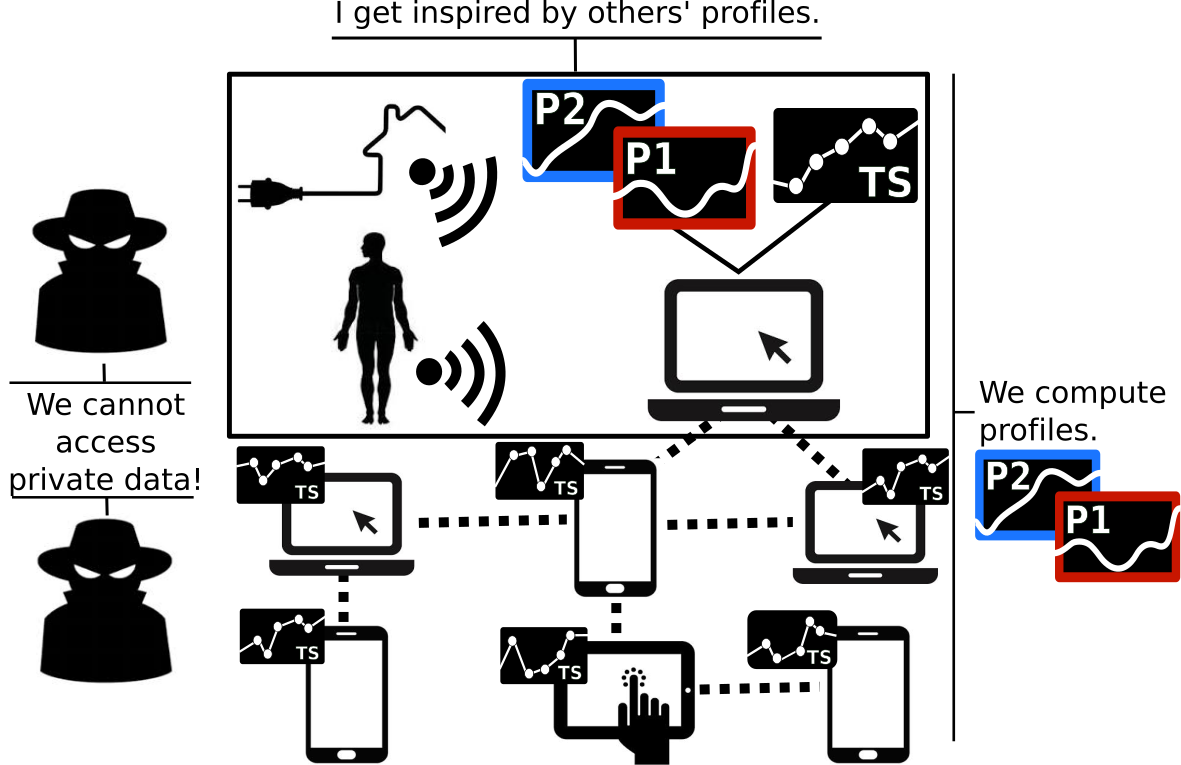
Chiaroscuro1 aims at:
- Extracting representative profiles
- From a set of time-series distributed over a large number of personal devices (massive populations)
- Without jeopardizing individuals' privacy
Chiaroscuro avoids two major common obstacles to personal data analytics.It does not copy the personal time-series on any central server (no information about a personal time-series leaves in a non-protected form its personal host devices). And it does not use any prohibitively-costly cryptographic protocol for extracting the profiles.
Chiaroscuro rather promotes an innovative approach that discloses perturbed agregate information all along the execution sequence by intertwining encryption schemes with sanitization schemes.
References
1. T. Allard, G. Hébrail, F. Masseglia, and E. Pacitti, "Chiaroscuro: Transparency and privacy for massive personal time-series clustering," in Proceedings of the 2015 ACM SIGMOD International Conference on Management of Data, SIGMOD ’15. ACM, 2015, pp. 779–794. Available here.